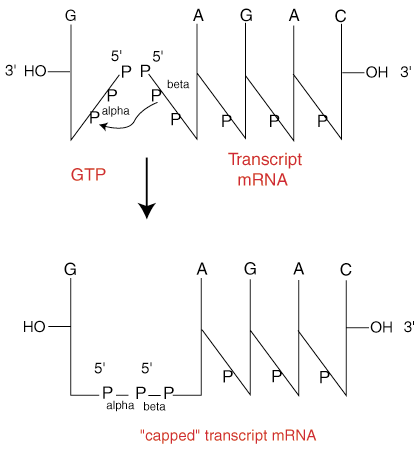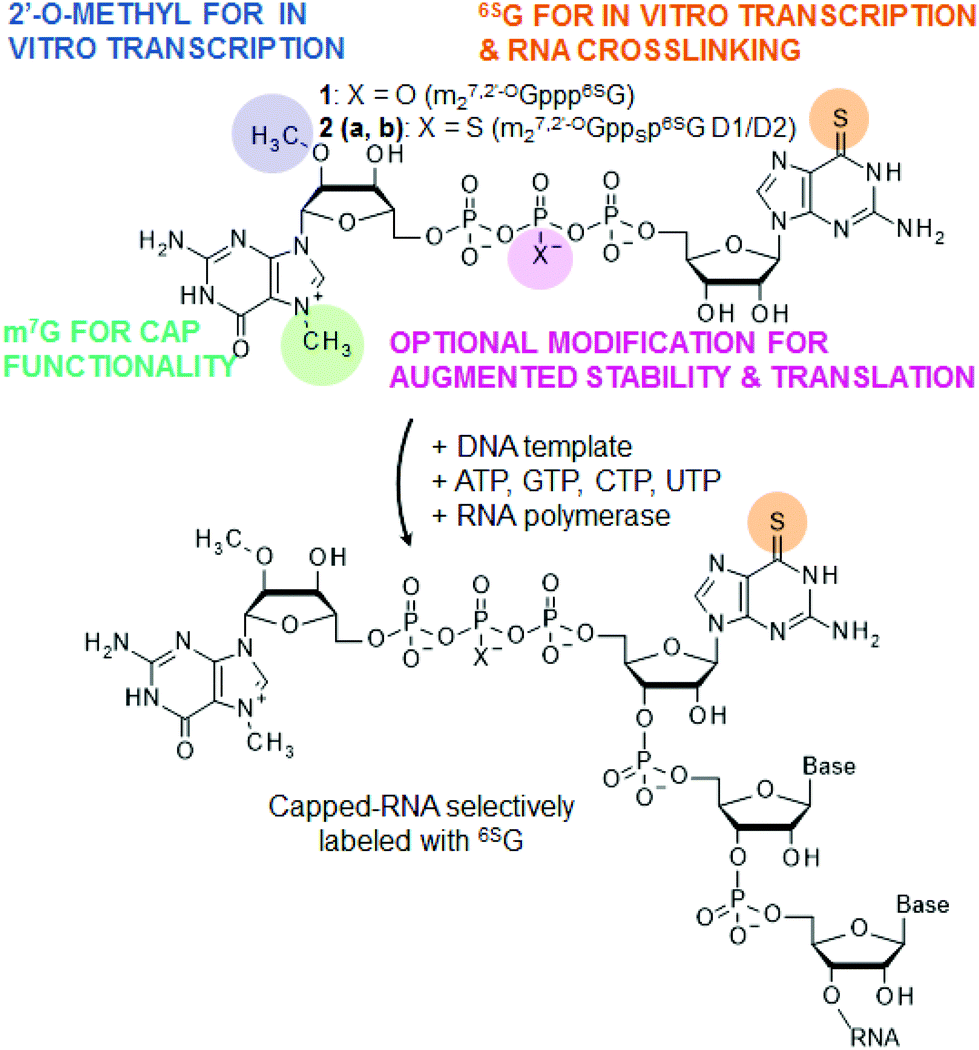
Cap analogs containing 6-thioguanosine – reagents for the synthesis of mRNAs selectively photo-crosslinkable with cap-binding biomolecules - Organic & Biomolecular Chemistry (RSC Publishing) DOI:10.1039/C4OB00059E

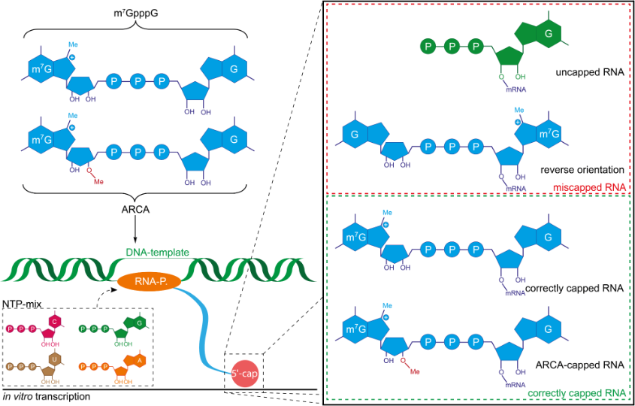
Schematic representation of co-transcriptional capping with different... | Download Scientific Diagram
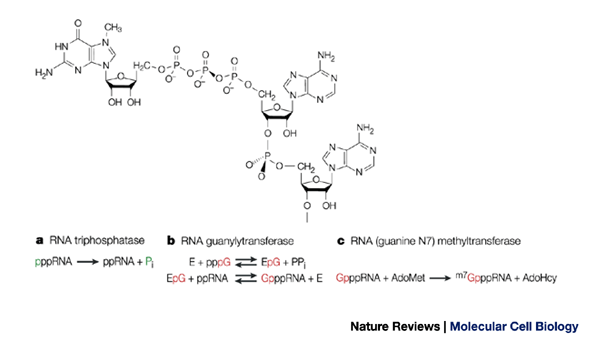
What messenger RNA capping tells us about eukaryotic evolution | Nature Reviews Molecular Cell Biology

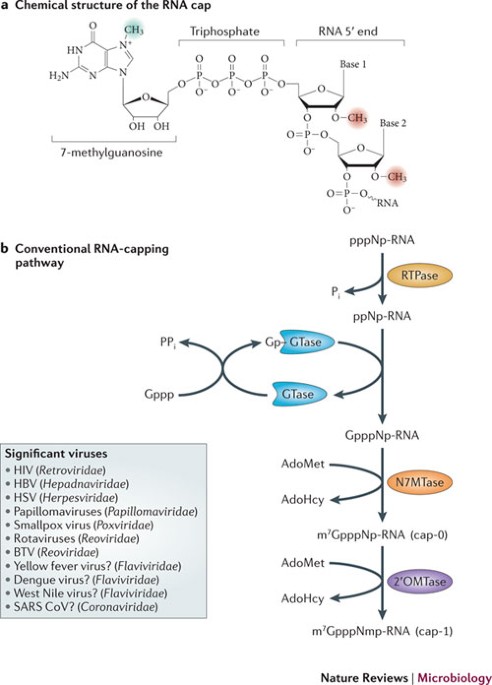
Capping mechanisms in RNA viruses. (A) The eukaryotic mRNA cap consists... | Download Scientific Diagram

RNA processing Contents 1 Capping 1.1 Features 1.2 Functions 1.3 Pathways of mRNA decay 1.4 Innate immunity and role of capping 1.4.1 Molecular basis of the recognition of 5'-ppp RNA by RIG-I 1.5 Synthetic cap analogs 2 Splicing Right after the ...

Mechanism that controls mRNA capping in gene regulation identified by Dundee interdisciplinary collaboration | School of Life Sciences

The expanding field of non-canonical RNA capping: new enzymes and mechanisms | Royal Society Open Science

CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology

A Mammalian Pre-mRNA 5′ End Capping Quality Control Mechanism and an Unexpected Link of Capping to Pre-mRNA Processing - ScienceDirect


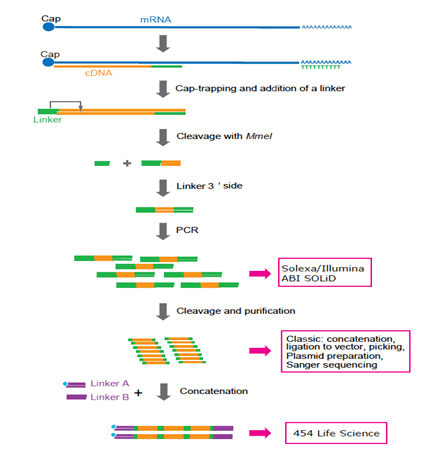




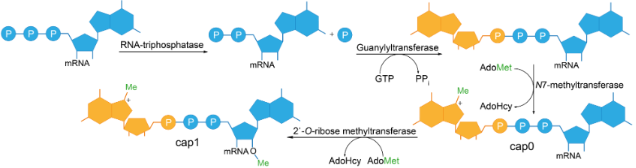
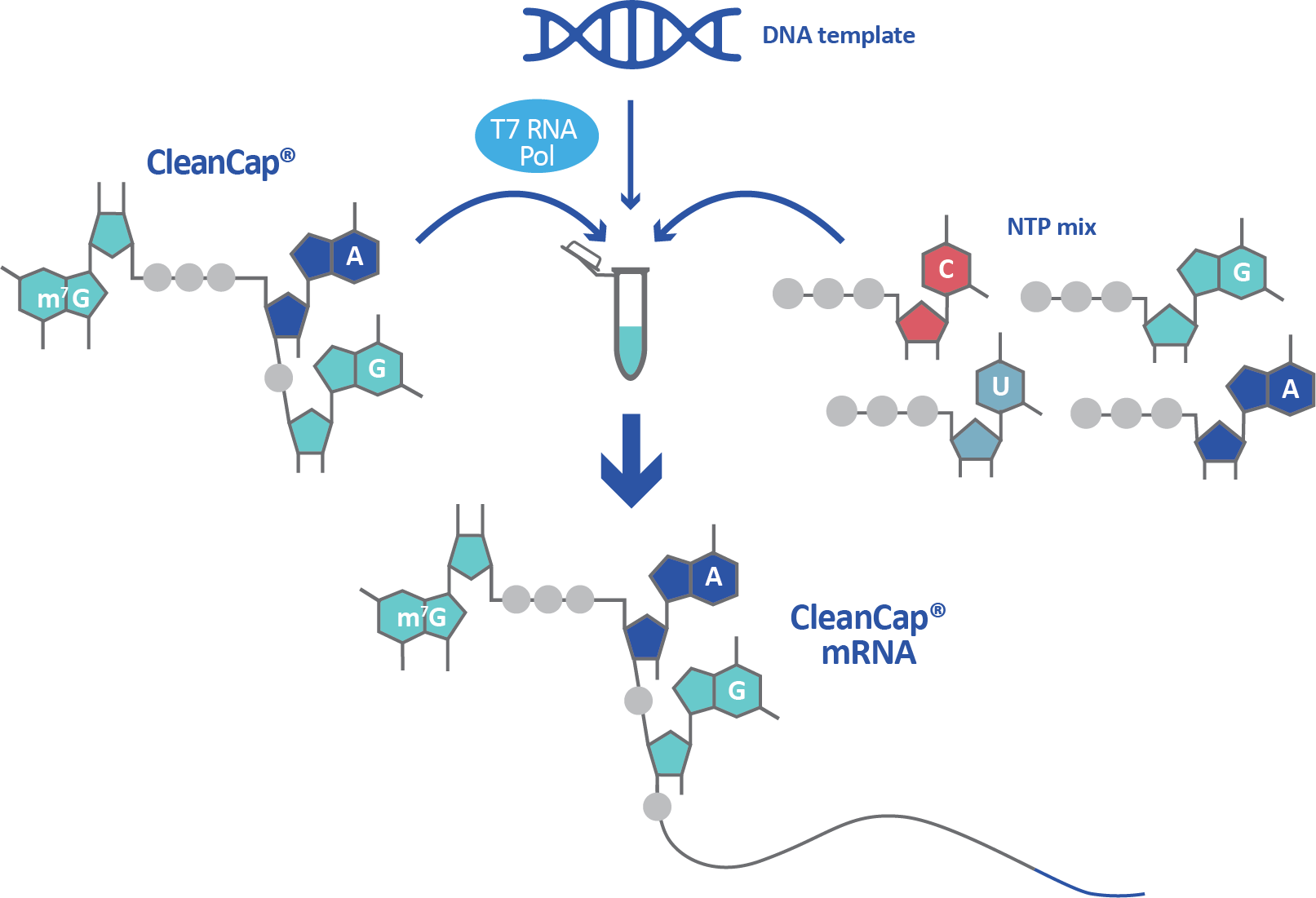
![PDF] Synthetic mRNA capping | Semantic Scholar PDF] Synthetic mRNA capping | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1e7ca15569c032d24a48d98f553afd31be586ad2/4-Figure3-1.png)