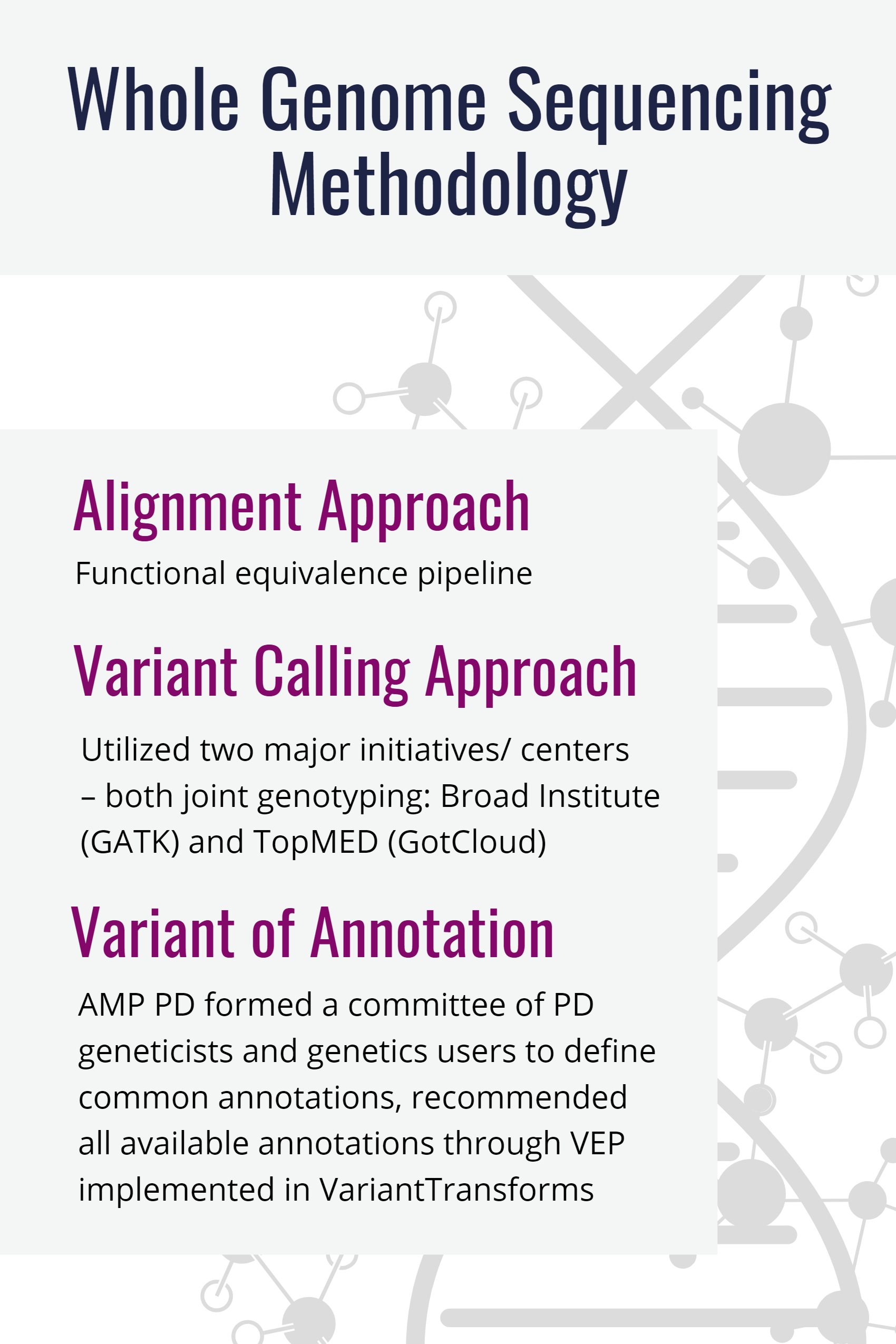
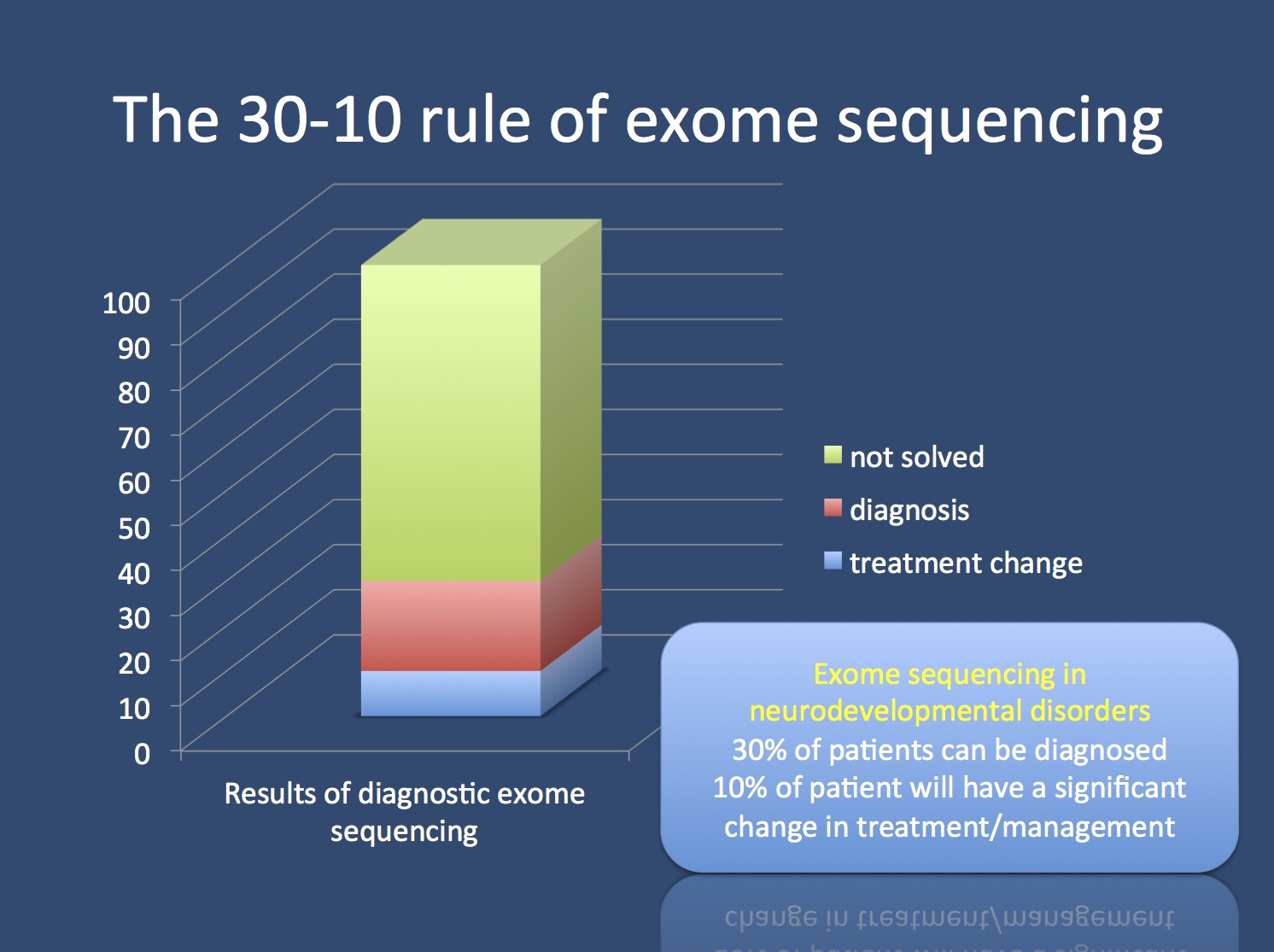
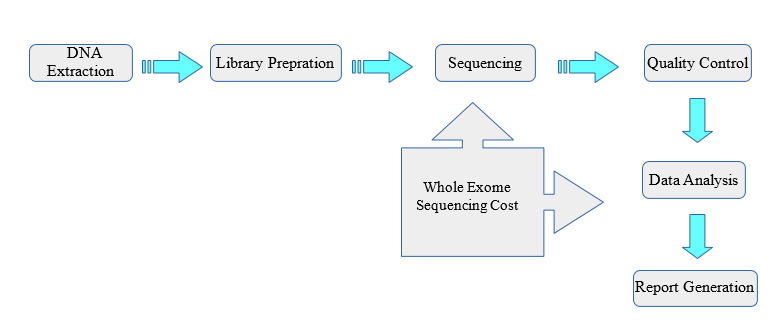
Pipeline of our study design. WES, whole exome sequencing; WGS, whole... | Download Scientific Diagram
![PDF] SeqAnt 2.0: Whole-Genome Annotation and Natural-Language Searching in the Cloud | Semantic Scholar PDF] SeqAnt 2.0: Whole-Genome Annotation and Natural-Language Searching in the Cloud | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/5a0c899dddf0e6814a11055aaeae28861a5f8a0a/4-Figure1-1.png)
PDF] SeqAnt 2.0: Whole-Genome Annotation and Natural-Language Searching in the Cloud | Semantic Scholar
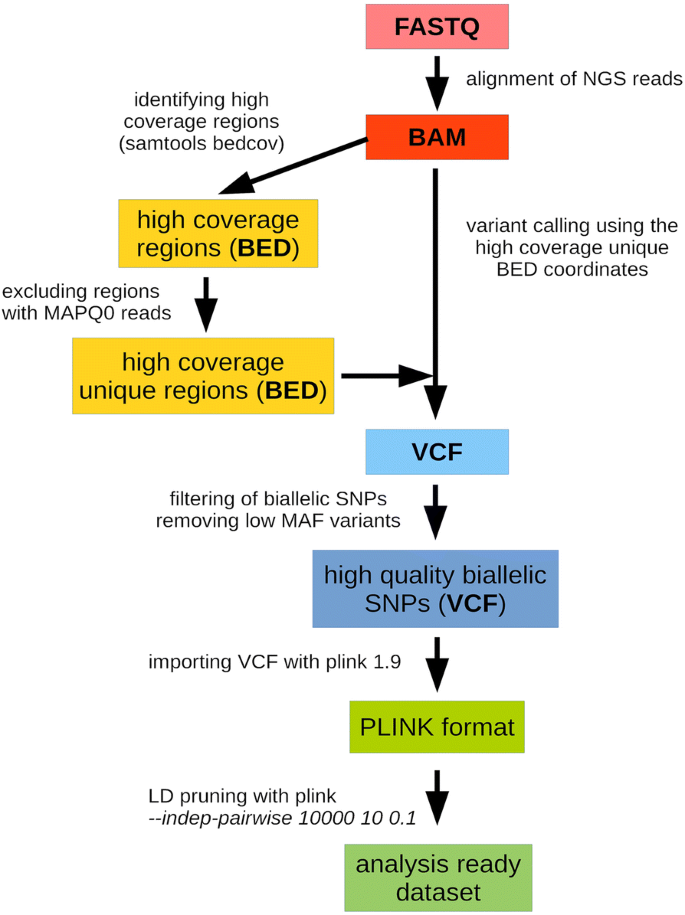
Evaluation of whole exome sequencing as an alternative to BeadChip and whole genome sequencing in human population genetic analysis | BMC Genomics | Full Text
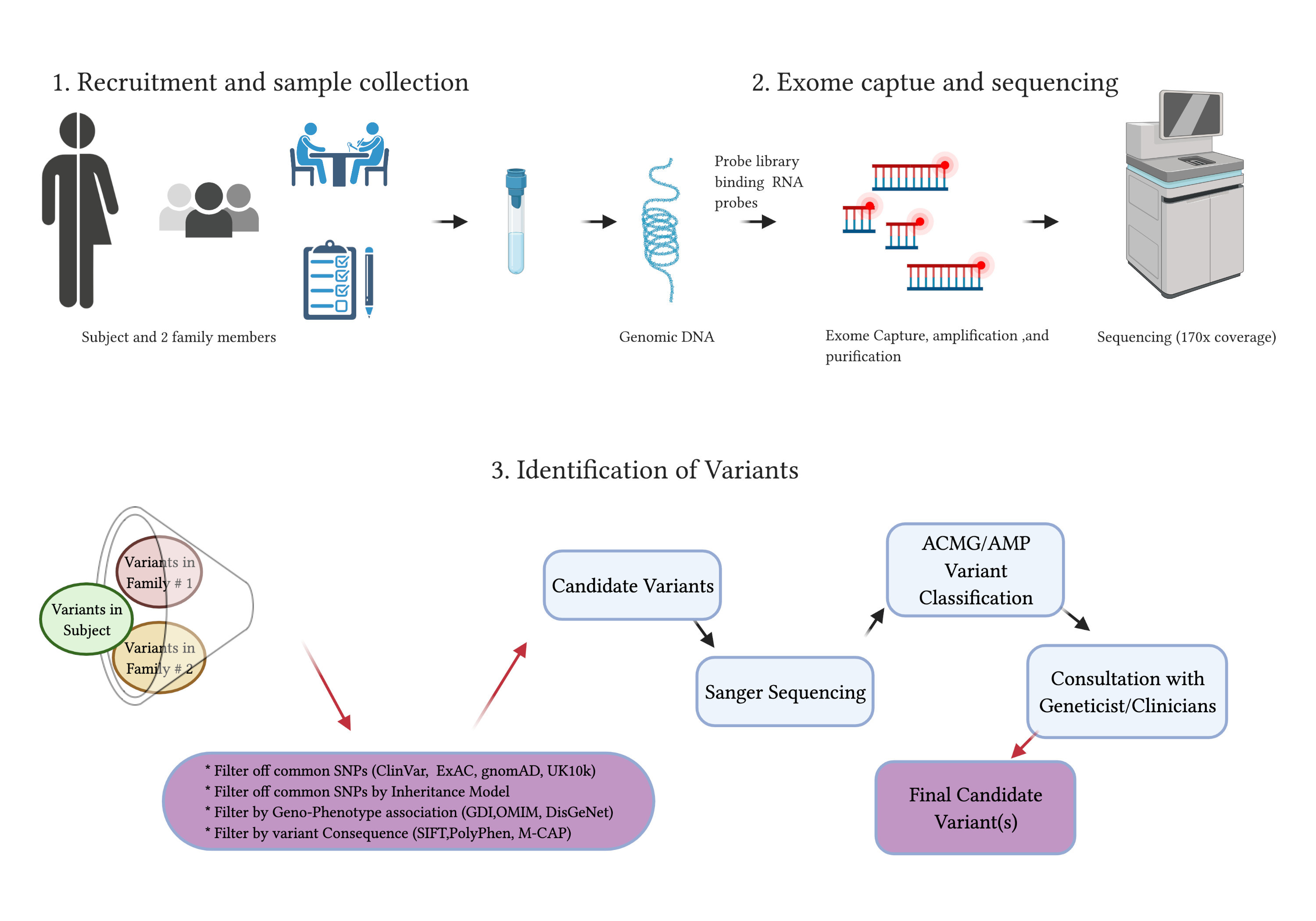
JCM | Free Full-Text | Whole-Exome Sequencing to Identify Potential Genetic Risk in Substance Use Disorders: A Pilot Feasibility Study | HTML

Overview of the whole exome sequencing study workflow. Abbreviations... | Download Scientific Diagram
1010Genome Presents Next Generation Sequencing and Whole Exome Sequencing Services | Quality NGS Bioinformatics Data Analysis Services

Modeling clonal structure over narrow time frames via circulating tumor DNA in metastatic breast cancer. - Abstract - Europe PMC

Overview of high-throughput emulsion whole-genome amplification. a The... | Download Scientific Diagram

Systematic dissection of biases in whole-exome and whole-genome sequencing reveals major determinants of coding sequence coverage | Scientific Reports

A 26-hour system of highly sensitive whole genome sequencing for emergency management of genetic diseases – topic of research paper in Biological sciences. Download scholarly article PDF and read for free on
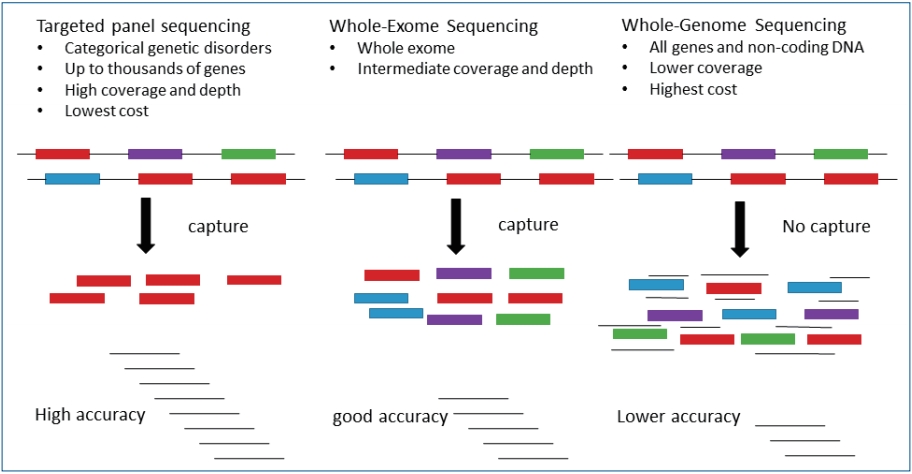
Genetic tests by next-generation sequencing in children with developmental delay and/or intellectual disability
PLOS ONE: The utility of Escherichia coli as a contamination indicator for rural drinking water: Evidence from whole genome sequencing
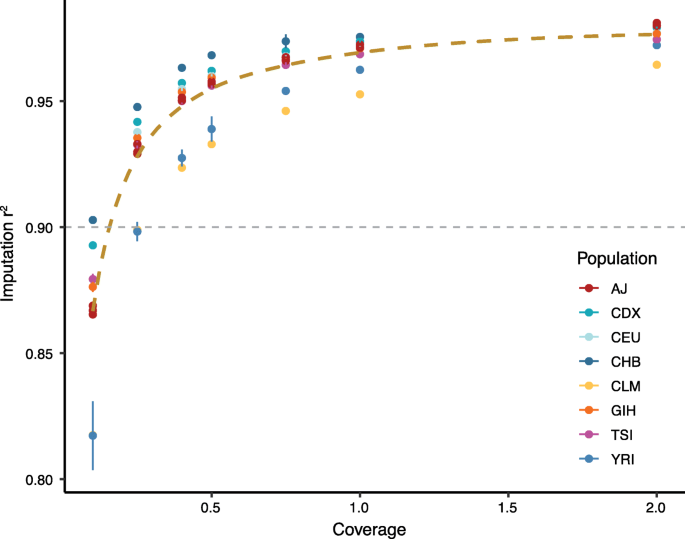
Low coverage whole genome sequencing enables accurate assessment of common variants and calculation of genome-wide polygenic scores | Genome Medicine | Full Text