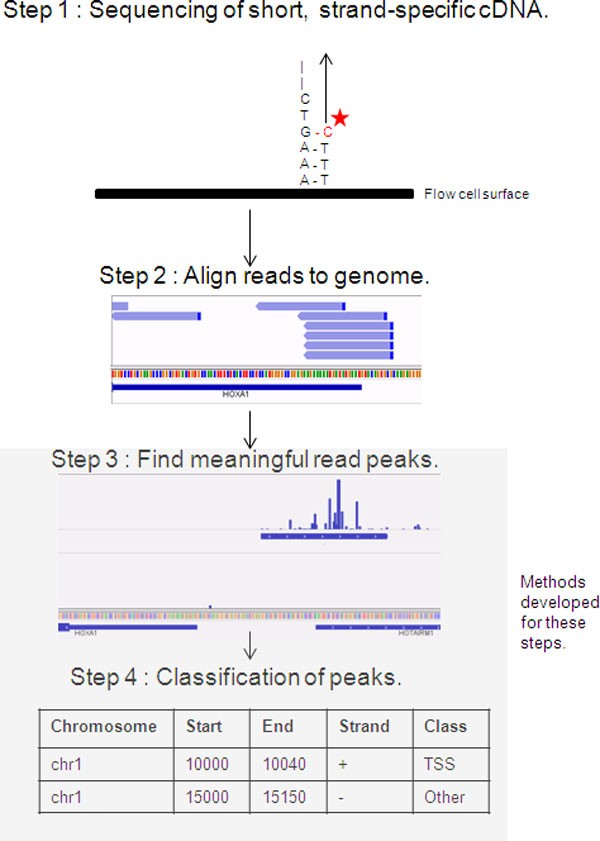
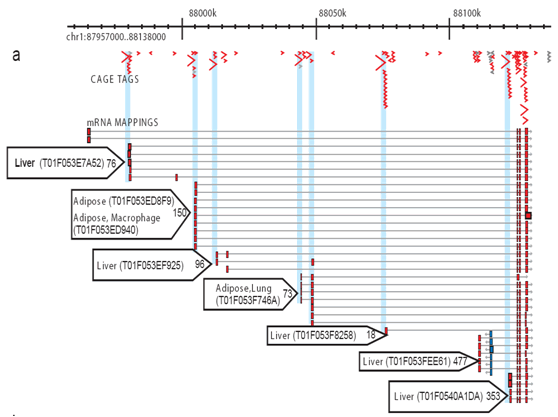
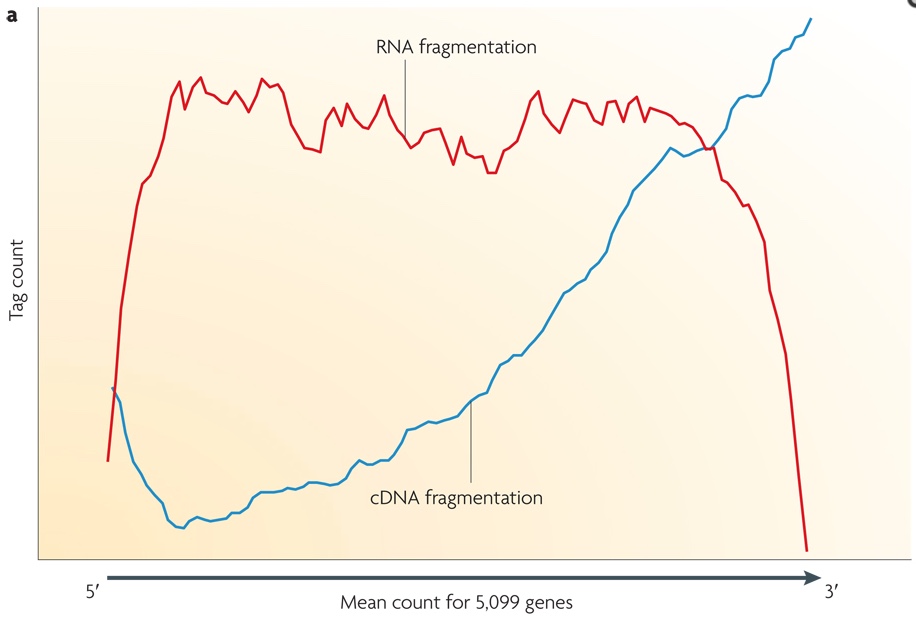
![PDF] Deep cap analysis gene expression (CAGE): genome-wide identification of promoters, quantification of their expression, and network inference. | Semantic Scholar PDF] Deep cap analysis gene expression (CAGE): genome-wide identification of promoters, quantification of their expression, and network inference. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6bbc59a3216052d585fdba17e64c4eea09933715/2-Table1-1.png)
PDF] Deep cap analysis gene expression (CAGE): genome-wide identification of promoters, quantification of their expression, and network inference. | Semantic Scholar

Workflow of the analysis pipeline and respective tools used for CAGE... | Download Scientific Diagram
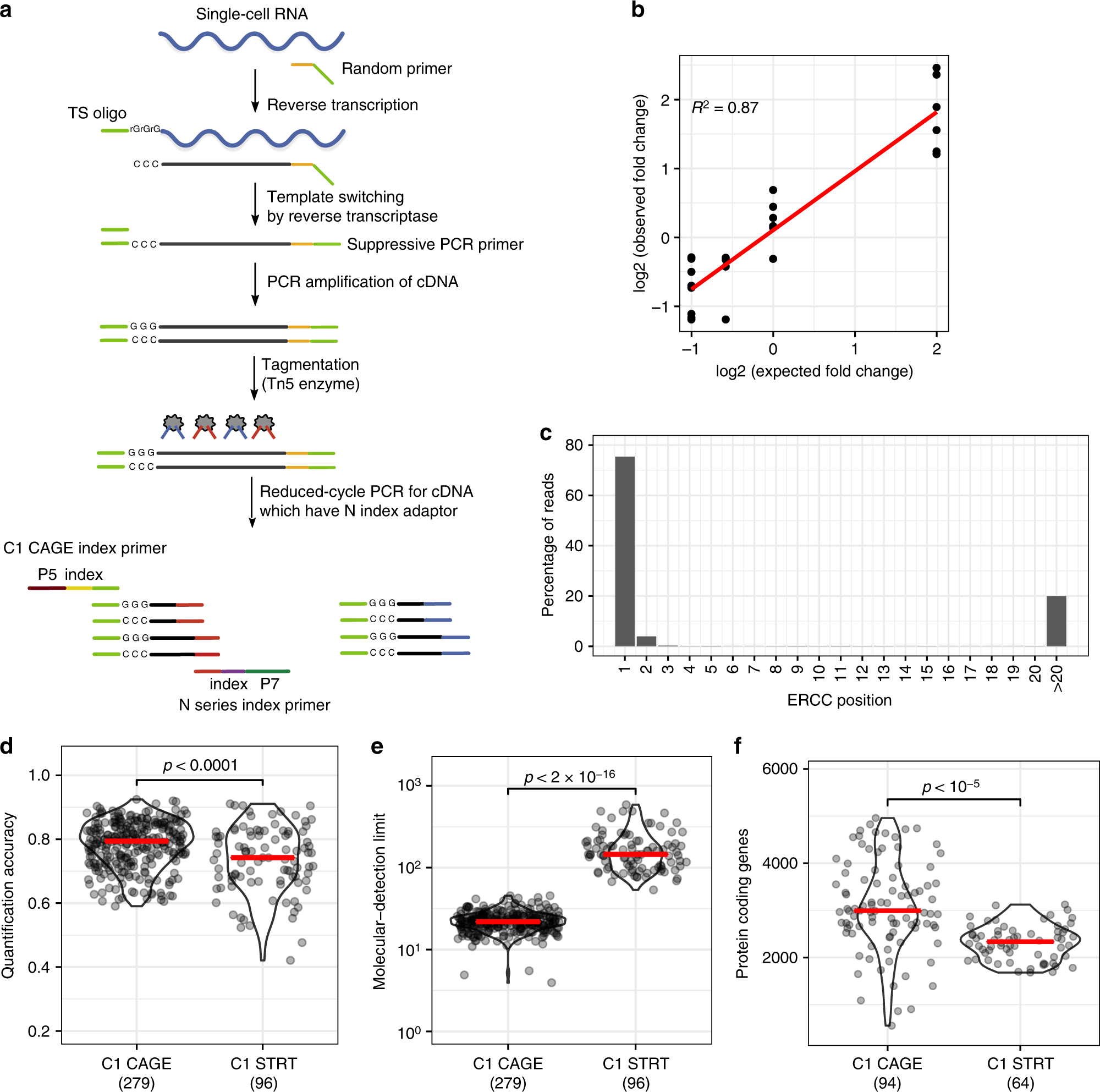
C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution | Nature Communications
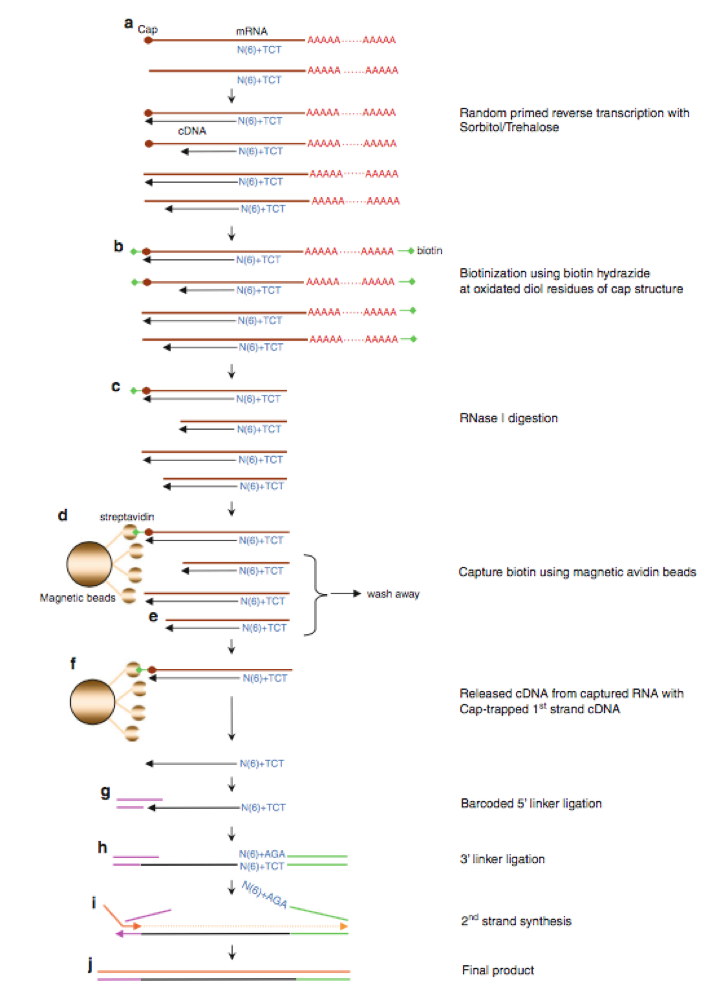
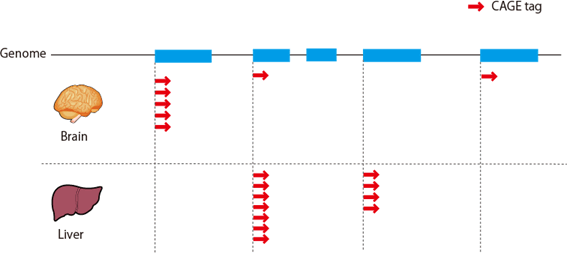
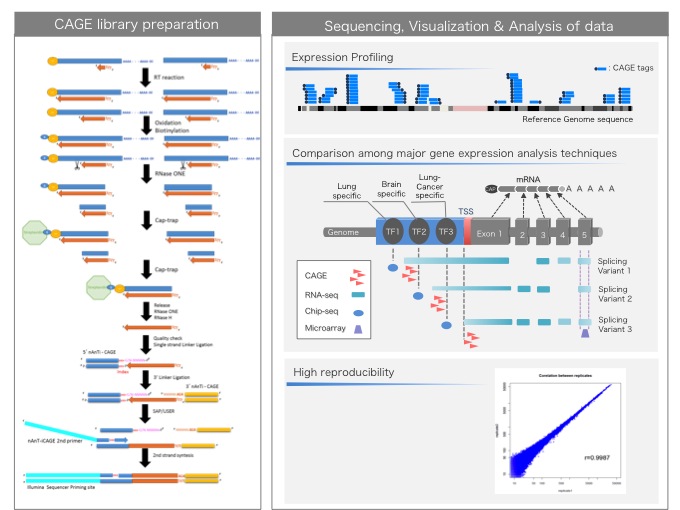
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター

Deep Cap Analysis of Gene Expression (CAGE): Genome-Wide Identification of Promoters, Quantification of Their Activity, and Transcriptional Network Inference | SpringerLink

DeepCAGE: Incorporating Transcription Factors in Genome-wide Prediction of Chromatin Accessibility | bioRxiv
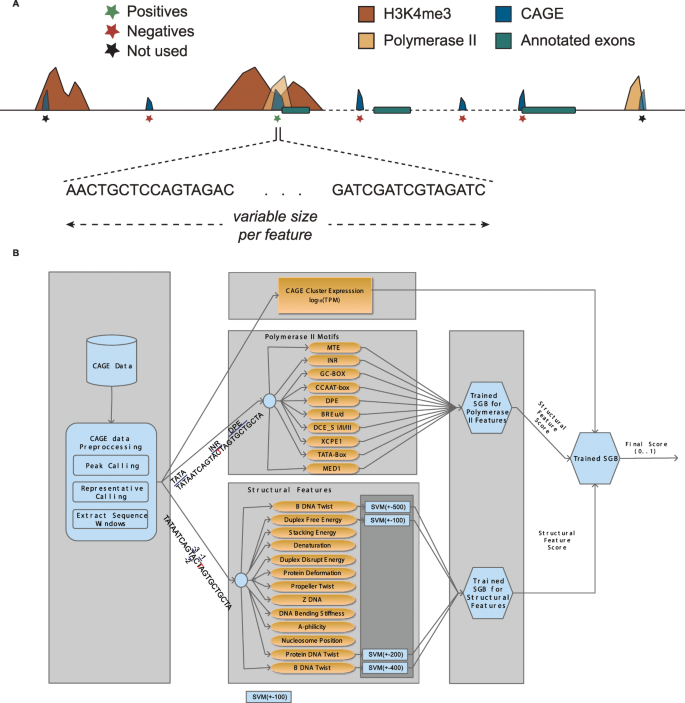
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports
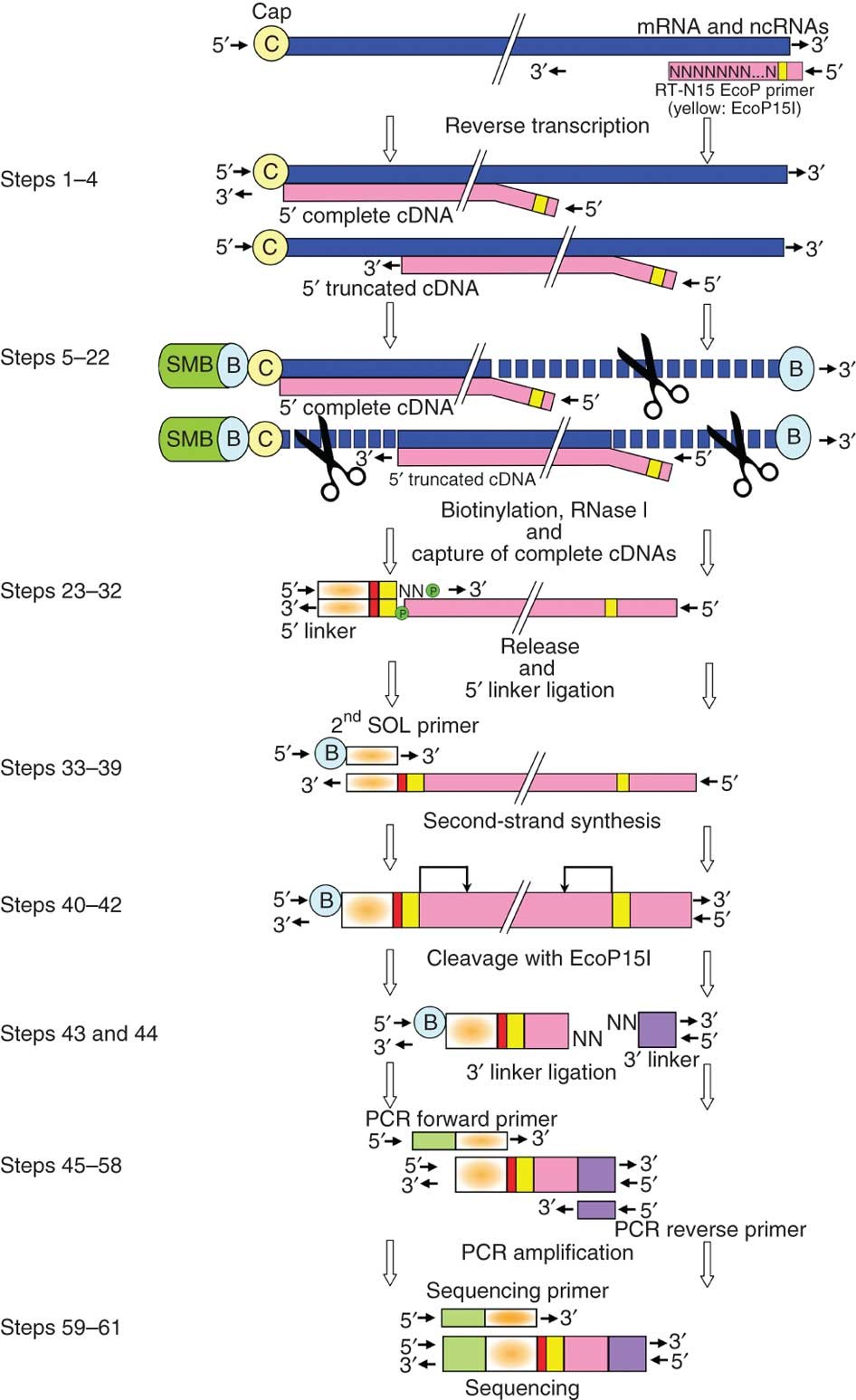
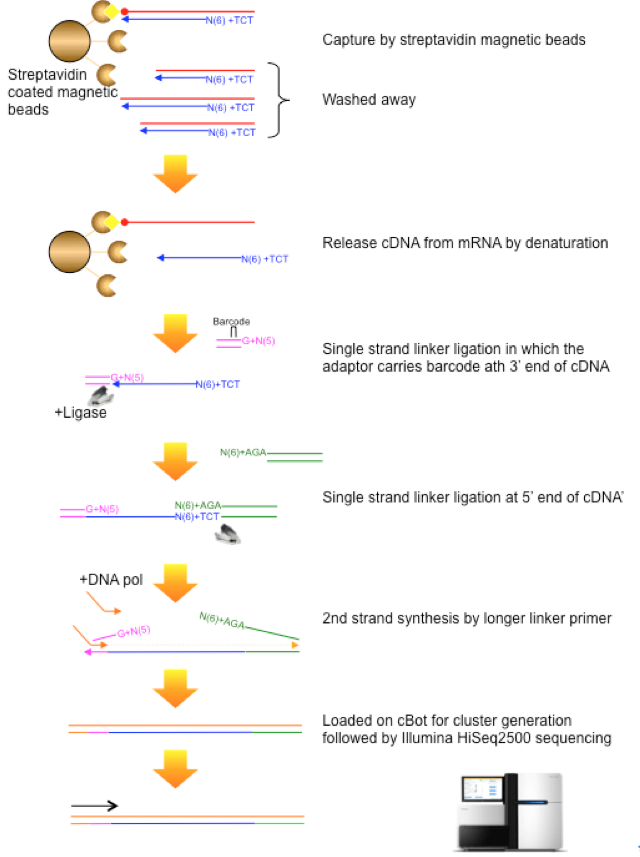
5′ end–centered expression profiling using cap-analysis gene expression and next-generation sequencing | Nature Protocols

Enhanced Identification of Transcriptional Enhancers Provides Mechanistic Insights into Diseases: Trends in Genetics

Deep Cap Analysis of Gene Expression (CAGE): Genome-Wide Identification of Promoters, Quantification of Their Activity, and Transcriptional Network Inference. | Semantic Scholar
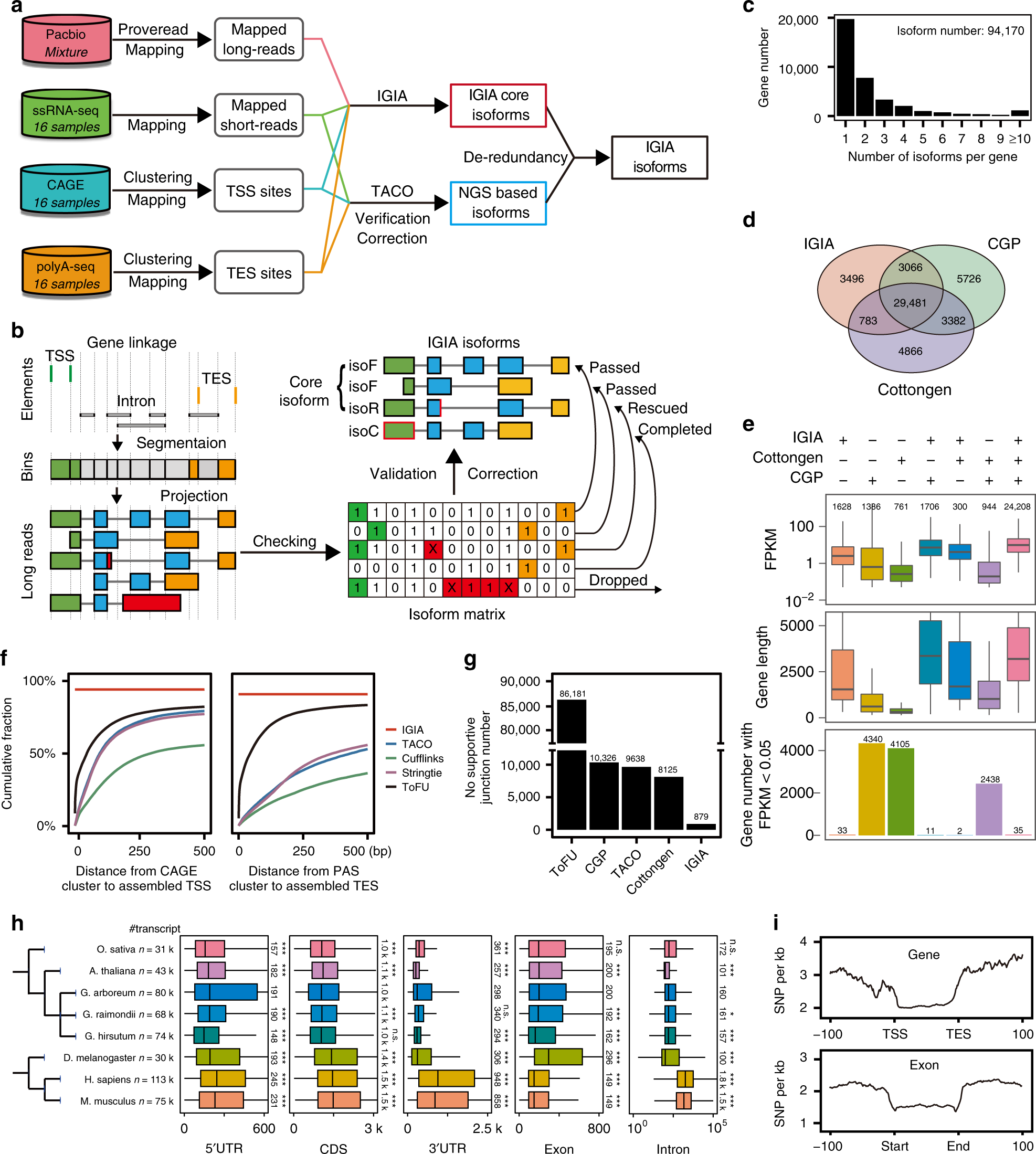
Multi-strategic RNA-seq analysis reveals a high-resolution transcriptional landscape in cotton | Nature Communications

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

piPipes – a set of pipelines for piRNA and transposon analysis via small RNA-seq, RNA-seq, degradome- and CAGE-seq, ChIP-seq and genomic DNA sequencing | RNA-Seq Blog